相结合)的正则化并没有产生更好的泛化表现。这可能与我们在介绍中提到的这个特定时间序列的特征有关。
plot(history, metrics = "loss")
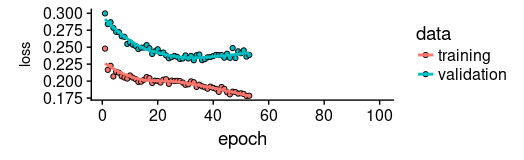
现在让我们看看该模型捕捉训练集特征的效果如何。
pred_train <- model %>%
predict(
X_train,
batch_size = FLAGS$batch_size) %>%
.[, , 1]
# Retransform values to original scale
pred_train <- (pred_train * scale_history + center_history) ^2
compare_train <- df %>%
filter(key == "training")
# build a dataframe that has both actual and predicted values
for (i in 1:nrow(pred_train))
{
varname <- paste0("pred_train", i)
compare_train <- mutate(
compare_train,
!!varname := c(
rep(NA, FLAGS$n_timesteps + i - 1),
pred_train[i,],
rep(NA,
nrow(compare_train) - FLAGS$n_timesteps * 2 - i + 1)))
}
我们计算所有预测序列的平均 RSME。
coln <- colnames(compare_train)[4:ncol(compare_train)]
cols <- map(coln, quo(sym(.)))
rsme_train <- map_dbl(
cols,
function(col)
{
rmse(
compare_train,
truth = value,
estimate = !!col,
na.rm = TRUE)
}) %>%
mean()
rsme_train
21.01495
这些预测看起来如何?由于所有预测序列的可视化看起来会非常拥挤,我们会间隔地选择起始点。
ggplot(
compare_train,
aes(x = index, y = value)) +
geom_line() +
geom_line(aes(y = pred_train1), color = "cyan") +
geom_line(aes(y = pred_train50), color = "red") +
geom_line(aes(y = pred_train100), color = "green") +
geom_line(aes(y = pred_train150), color = "violet") +
geom_line(aes(y = pred_train200), color = "cyan") +
geom_line(aes(y = pred_train250), color = "red") +
geom_line(aes(y = pred_train300), color = "red") +
geom_line(aes(y = pred_train350), color = "green") +
geom_line(aes(y = pred_train400), color = "cyan") +
geom_line(aes(y = pred_train450), color = "red") +
geom_line(aes(y = pred_train500), color = "green") +
geom_line(aes(y = pred_train550), color = "violet") +
geom_line(aes(y = pred_train600), color = "cyan") +
geom_line(aes(y = pred_train650), color = "red") +
geom_line(aes(y = pred_train700), color = "red") +
geom_line(aes(y = pred_train750), color = "green") +
ggtitle("Predictions on the training set")
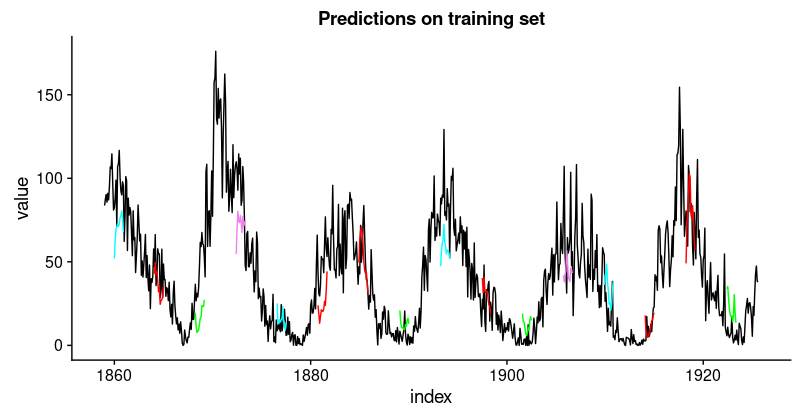
看起来像当好。对于验证集,我们并不认为有同样好的结果。
pred_test <- model %>%
predict(
X_test,
batch_size = FLAGS$batch_size) %>%
.[, , 1]
# Retransform values to original scale
pred_test <- (pred_test * scale_history + center_history) ^2
pred_test[1:10, 1:5] %>% print()
compare_test <- df %>% filter(key == "testing")
# build a dataframe that has both actual and predicted values
for (i in 1:nrow(pred_test))
{
varname <- paste0("pred_test", i)
compare_test <-mutate(
compare_test,
!!varname := c(
rep(NA, FLAGS$n_timesteps + i - 1),
pred_test[i,],
rep(NA,
nrow(compare_test) - FLAGS$n_timesteps * 2 - i + 1)))
}
compare_test %>% write_csv(
str_replace(model_path, ".hdf5", ".test.csv"))
compare_test[FLAGS$n_timesteps:(FLAGS$n_timesteps + 10), c(2, 4:8)] %>%
print()
coln <- colnames(compare_test)[4:ncol(compare_test)]
cols <- map(coln, quo(sym(.)))
rsme_test <- map_dbl(
cols,
function(col)
{
rmse(
compare_test,
truth = value,
estimate = !!col,
na.rm = TRUE)
}) %>%
mean()
rsme_test
31.31616
ggplot(
compare_test,
aes(x = index, y = value)) +
geom_line() +
geom_line(aes(y = pred_test